Research
My research interest is in bioinformatics. I use computer science techniques and emerging technologies to develop accurate and scalable bioinformatics tools. Currently, the two main areas of my research are clinical genomics and agroinformatics.
Clinical genomics
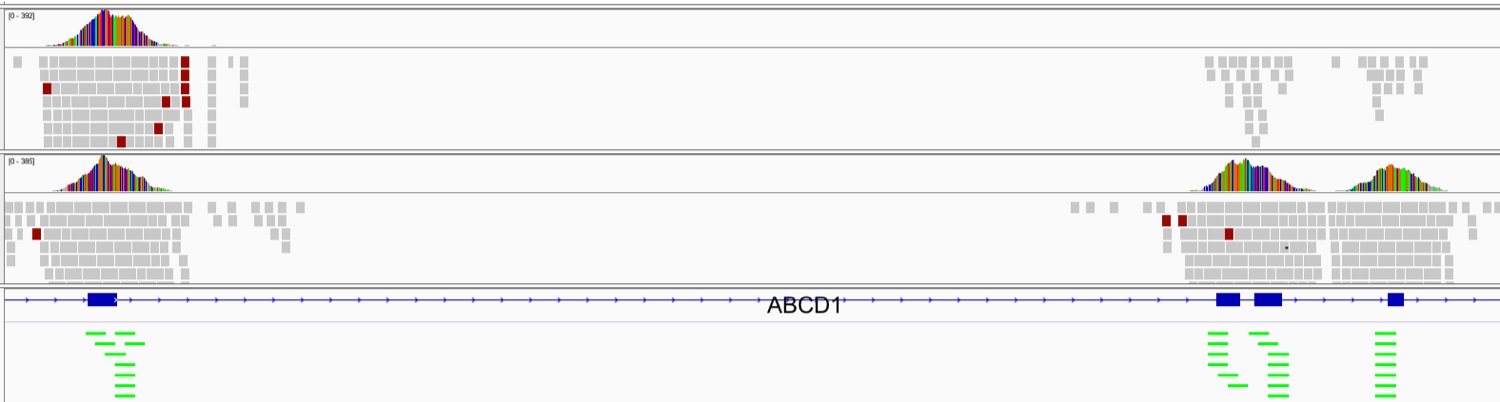
Next-Generation Sequencing (NGS) has shown great promise in aiding clinical diagnosis of genetic disorders, especially for disorders where multiple genes are implicated. Copy Number Variations (CNVs) are known to cause genetic disorders. Detecting clinically relevant CNVs is still a challenge. Among other limitations, existing tools for detecting CNVs still suffer from a high rates of false positives. My work in this area focuses on developing methods and tools to detect clinically relevant CNVs on whole-exome and whole-genome data.
Agro-Informatics
Food insecurity continues to be a problem around the world, including in developed nations. To help tackle this problem, groups such as GEMS are working on platforms that will enable integrative analysis of genotype, environmental and management data. Collectively, this data could be used for data-driven plant breeding efforts. In collaboration with GEMS , I am working on techniques for integrative analysis of genotype, environmental and management data.